jnmaciuch
Senior Member (Voting Rights)
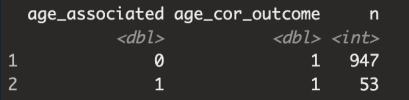
But that's the exact scenario i'm flagging. That's what they said in the paper. There was no association between all the other NII-whatever variables (besides NII-RF) when tested individually, but they did become significant when a covariate was added. So what you're simulating here is not the issue I'm bringing up.This is because in the model you made, there is no relationship between ME/CFS and niXX, so it is all due to chance. niXX is just a function of age, so there should only be about 5% significant as false positives when testing the association with mecfs_status.
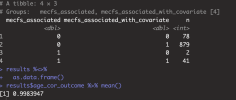
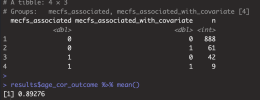
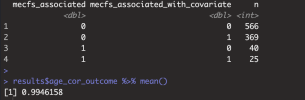
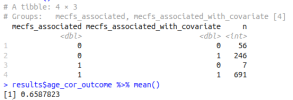
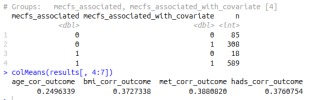
If niXX actually depends on mecfs_status, then adding the covariate can make it more significant, and can take p from greater than 0.05 to less than 0.05. I added mecfs_status to the initial niXX model in your code:
I'm saying that this is likely human error because you can't force an initially unassociated variable to become significant with a covariate (more than by chance) even if the covariates are significantly associated with outcome.