I expect @SNT Gatchaman is busy on other things right now, but I hope he will comment more on the paper at some point.
You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
Evidence of White Matter Neuroinflammation in [ME/CFS]: A Diffusion-Based Neuroinflammation Imaging Study 2026 Yu et al
jnmaciuch
Senior Member (Voting Rights)
(Int = fitted model intercept)
Testing the association of NII-RF and ME/CFS without covariates:
p-value of ME/CFS derived by comparing difference in model performance between:
model1: NII-RF ~ Int
model2: NII-RF ~ Int + Beta1 * ME/CFS (0/1 binary)
model1 and model2 each have their own precision. If a model containing ME/CFS is better at predicting NII-RF scores than a model containing just the intercept, we say there is a significant association.
Testing the association of NII-RF and ME/CFS with covariates:
p-value of ME/CFS derived by comparing difference in model performance between:
model3: NII-RF ~ Int + Beta1 * age
model4: NII-RF ~ Int + Beta1 * age + Beta2 * ME/CFS (0/1 binary)
If the model performance improves when the ME/CFS variable is added, we say that ME/CFS is significantly associated with NII-RF accounting for age.
If ME/CFS has any predictive power for NII-RF, you will get p < 0.05 comparing models 1 and 2. If you want to ask whether that association is confounded by age, you compare models 3 and 4. The improvement in precision gained by adding age would be present in both models, so it doesn't matter.
[Edit: and for ME/CFS to be significant in the model3 vs. model4 comparison, it by definition has to have good predictive power for NII-RF. So you see why it would be weird to get significance comparing models 3 and 4, but no significance comparing models 1 and 2 (for all the measurements except NII-RF, which was significant in both comparisons)]
Testing the association of NII-RF and ME/CFS without covariates:
p-value of ME/CFS derived by comparing difference in model performance between:
model1: NII-RF ~ Int
model2: NII-RF ~ Int + Beta1 * ME/CFS (0/1 binary)
model1 and model2 each have their own precision. If a model containing ME/CFS is better at predicting NII-RF scores than a model containing just the intercept, we say there is a significant association.
Testing the association of NII-RF and ME/CFS with covariates:
p-value of ME/CFS derived by comparing difference in model performance between:
model3: NII-RF ~ Int + Beta1 * age
model4: NII-RF ~ Int + Beta1 * age + Beta2 * ME/CFS (0/1 binary)
If the model performance improves when the ME/CFS variable is added, we say that ME/CFS is significantly associated with NII-RF accounting for age.
If ME/CFS has any predictive power for NII-RF, you will get p < 0.05 comparing models 1 and 2. If you want to ask whether that association is confounded by age, you compare models 3 and 4. The improvement in precision gained by adding age would be present in both models, so it doesn't matter.
[Edit: and for ME/CFS to be significant in the model3 vs. model4 comparison, it by definition has to have good predictive power for NII-RF. So you see why it would be weird to get significance comparing models 3 and 4, but no significance comparing models 1 and 2 (for all the measurements except NII-RF, which was significant in both comparisons)]
Last edited:
In the case of models 3 and 4, we can be more confident about the amount that ME/CFS status improves the model, since the age variable is included and can explain some of the variance.The improvement in precision gained by adding age would be present in both models, so it doesn't matter.
I'm not able to right now, but I might code it with simulated data later. It should be relatively straightforward to test what happens in this scenario when NII-RF is correlated to both age and ME/CFS, but where age and ME/CFS are not correlated to each other.
jnmaciuch
Senior Member (Voting Rights)
That’s what I’ve been trying to explain this whole time. It simply does not matter. The way the comparison is structured, it’s only looking at the difference between two models where the only change is whether ME/CFS is added to the model.It should be relatively straightforward to test what happens in this scenario when NII-RF is correlated to both age and ME/CFS, but where age and ME/CFS are not correlated to each other.
It does not matter at all if age has a separate association with NII-RF because the p-value theyre reporting should be from testing whether a model with age and ME/CFS performs significantly better than a model with age alone. Any performance boost from age is kept constant.
Jonathan Edwards
Senior Member (Voting Rights)
The pictures are interesting in that the colour signals are mostly in sensory association areas and seem biased to the right. The pattern does not look like an inflammatory one to me unless there is an immune reaction to a local antigen associated with particular functions. Even then, the bias to right looks much more like a neurocognitive pattern than an immunological one.
I wonder if we are just looking at effects of (shifts in water associated with) blood flow associated with neural function - rather as in BOLD studies.
It is a pity that the authors keep referring to neuroinflammation as if it was a homogeneous entity rather than sticking to shifts in physiology.
I wonder if we are just looking at effects of (shifts in water associated with) blood flow associated with neural function - rather as in BOLD studies.
It is a pity that the authors keep referring to neuroinflammation as if it was a homogeneous entity rather than sticking to shifts in physiology.
jnmaciuch
Senior Member (Voting Rights)
The math couldnt work out like that anyways, that’s what I’m trying to explain in my conversation with forestglip. You’re supposed to start with the univariate analysis, seeing if ME/CFS on its own is associated. If it is, then you control for confounders. The outcome will be that either the association is still significant (meaning that the confounders don’t matter) or it’s no longer significant (meaning that the first association is explained by the confounder). To get no association in the univariate analysis but a significant association in the multivariate means that something very funky happened in one of the calculations (as in human error)Yes, I had the same concern about the addition of controlling for the 'confounders' producing significant results that would not have been there without. I think the age and sex probably are valid, but I thought that the groups were matched on those anyway, in which case there is less reason to do that.
It’s like if you set up an experiment to test whether time of day matters in how you perform at ring toss. Your hypothesis is that you get more tired as the day goes on so your performance will decrease later in the day.
You test it out at 3 different times of day, and see that your scores are consistent no matter the time. Despite already finding no association, you decide to test if amount of light explains the “association” between time of day and ring toss performance. So you test your ring toss at the same 3 times of day, but for each timepoint you do a test with the windows open and with the windows drawn shut. By some weird happenstance, you managed to score every toss you made at the latest time point, even when you were in complete darkness. In your publication, you report that time of day significantly affects ring tossing ability and that this association is not explained by amount of light.
That’s basically what that part of the paper reads like to me.
It would be nice to see charts like that one in the Keri paper of the actual signals, with some indication of variability. I wonder if it is possible to average the signals for specific tissues for each cohort and present them as a chart like the Keri chart?For some reason the chart I attached upthread isn't showing (fixed now) - here it is again, from the Keri paper:
A question from ignorance: I don't know to what extent blood vessels (or indeed temporary increased blood flow) contribute to those isotropic signals? Maybe increased vascularity or blood vessel tortuosity or blood vessels that are over-leaky or just blood flow might increase the free fraction and so reduce the restricted fraction?
I'm not clear on how these isotropic signals are labelled relative fractions e.g. RF is the restricted fraction, but why the HR is called the Hindered Ratio, and seems to be the hindered signal relative to the total isotropic signal. I would have thought a relative fraction is already a ratio.
TY, I'm hoping this w/e will give sufficient time to look properly through the foundational techniques papers to try and make sense of the problems being discussed above. On my initial read of the paper one thing that always raises an eyebrow for me with these types of findings is the asymmetry. Eg —I expect @SNT Gatchaman is busy on other things right now, but I hope he will comment more on the paper at some point.
Similarly, a negative association between NII-FF and disease severity suggests that reduced apparent axonal density in the right superior longitudinal fasciculus and right posterior corona radiata may accompany worsening clinical presentation.
on average, patients show higher NII-FF, but the most severely affected show NII-FF loss in specific tracts. One tract (right posterior corona radiata) also showed group level NII-FF increases, implying that NII-FF may initially rise (e.g., reduced dispersion or compartmental re-weighting) and later fall with more severe disease (axon or packing loss), a pattern compatible with stage dependent pathology.
I don't see our pathology ending up being lateralised like this. ME/CFS has a very specific core symptom set, albeit with +/- other symptoms. A disease like MS can have foci in discrete and more or less random brain and spine regions leading to predictable motor/sensory/cognitive deficits. If ME/CFS occurs due to something like a a perfusion or metabolic vulnerability or some immune-driven process then what is the disease where it hits say the left superior longitudinal fasciculus and left posterior corona radiata?
Even once I have had a chance to get my head around the background of this newer technique, I suspect I won't have much additional to offer. Thank you to @Hutan for asking for clarification from the authors, and the analysis of the statistical reporting above may well be the more important discussion points.
However, all these newer neuroimaging techniques do seem like forward progress for us overall. Another new technique (paywalled paper so I haven't posted separately) that might be good for us is
A reference-based PET/MRI method for quantifying activation-induced changes in cerebral oxygen metabolism (2025, Journal of Cerebral Blood Flow & Metabolism)
Hybrid PET/MRI can overcome the complexity of PET imaging of the cerebral metabolic rate of oxygen (CMRO2), while retaining the ability to directly measure oxygen uptake in the brain. One technique, PMROx, incorporates complementary MRI methods acquired simultaneously with [15O]O2-PET. Specifically, the MRI-based method arterial spin labelling (ASL) to image cerebral blood flow (CBF) and MR-susceptometry to measure whole-brain CMRO2. PMROx is non-invasive with imaging times around 5 min, making it feasible to image CMRO2 under different conditions in one session.
This study presents the first application of PMROx to humans with the aims of evaluating its reliability and sensitivity to increased CMRO2 during functional activation (right-handed sequential finger tapping). In addition, blood-oxygen level dependent (BOLD) images were acquired to compare BOLD contrast to underlying changes in CBF and CMRO2. Across 14 participants, mean CMRO2 was 3.2 ± 0.5 mLO2/100 g/min with excellent within-session repeatability. Significant increases in CBF, CMRO2 and BOLD contrast were detected in the primary sensorimotor cortex, supplementary motor area and secondary somatosensory cortex.
This study demonstrated the ability of PMROx to image CMRO2 non-invasively, its sensitivity to increased regional CMRO2 and how PET/MRI provides the opportunity to compare BOLD contrast to underlying changes in perfusion and oxygen metabolism.
This study presents the first application of PMROx to humans with the aims of evaluating its reliability and sensitivity to increased CMRO2 during functional activation (right-handed sequential finger tapping). In addition, blood-oxygen level dependent (BOLD) images were acquired to compare BOLD contrast to underlying changes in CBF and CMRO2. Across 14 participants, mean CMRO2 was 3.2 ± 0.5 mLO2/100 g/min with excellent within-session repeatability. Significant increases in CBF, CMRO2 and BOLD contrast were detected in the primary sensorimotor cortex, supplementary motor area and secondary somatosensory cortex.
This study demonstrated the ability of PMROx to image CMRO2 non-invasively, its sensitivity to increased regional CMRO2 and how PET/MRI provides the opportunity to compare BOLD contrast to underlying changes in perfusion and oxygen metabolism.
I'm not sure about that.The math couldnt work out like that anyways, that’s what I’m trying to explain in my conversation with forestglip. You’re supposed to start with the univariate analysis, seeing if ME/CFS on its own is associated. If it is, then you control for confounders. The outcome will be that either the association is still significant (meaning that the confounders don’t matter) or it’s no longer significant (meaning that the first association is explained by the confounder). To get no association in the univariate analysis but a significant association in the multivariate means that something very funky happened in one of the calculations (as in human error)
Say you have a feature that increases a lot by age but is reliably lower in ME/CFS at each age. If you had a badly matched control group, a lot younger, then there might not be a significant difference between the ME/CFS group and the control group for that feature. So, there would be no obvious association to start with. But then, if you did control for age, the difference would appear.
jnmaciuch
Senior Member (Voting Rights)
It would not because of what I explained in post #69. When you report the p-value for a variable in a model with added covariates, you are reporting how well a model with ME/CFS plus all the covariates predicts the outcome variable compared to a model with all the covariates. That's what it means to "control for confounders" in a regression analysisSay you have a feature that increases a lot by age but is reliably lower in ME/CFS at each age. If you had a badly matched control group, a lot younger, then there might not be a significant difference between the ME/CFS group and the control group for that feature. So, there would be no obvious association to start with. But then, if you did control for age, the difference would appear.
Here is some R code simulating this with random data where NII-RF is correlated to age and to ME/CFS status:(Int = fitted model intercept)
Testing the association of NII-RF and ME/CFS without covariates:
p-value of ME/CFS derived by comparing difference in model performance between:
model1: NII-RF ~ Int
model2: NII-RF ~ Int + Beta1 * ME/CFS (0/1 binary)
model1 and model2 each have their own precision. If a model containing ME/CFS is better at predicting NII-RF scores than a model containing just the intercept, we say there is a significant association.
Testing the association of NII-RF and ME/CFS with covariates:
p-value of ME/CFS derived by comparing difference in model performance between:
model3: NII-RF ~ Int + Beta1 * age
model4: NII-RF ~ Int + Beta1 * age + Beta2 * ME/CFS (0/1 binary)
If the model performance improves when the ME/CFS variable is added, we say that ME/CFS is significantly associated with NII-RF accounting for age.
If ME/CFS has any predictive power for NII-RF, you will get p < 0.05 comparing models 1 and 2. If you want to ask whether that association is confounded by age, you compare models 3 and 4. The improvement in precision gained by adding age would be present in both models, so it doesn't matter.
[Edit: and for ME/CFS to be significant in the model3 vs. model4 comparison, it by definition has to have good predictive power for NII-RF. So you see why it would be weird to get significance comparing models 3 and 4, but no significance comparing models 1 and 2 (for all the measurements except NII-RF, which was significant in both comparisons)]
Code:
total_trials <- 1000
more_significant_count <- 0
for (i in 1:total_trials) {
age <- rnorm(50, mean = 45, sd = 10)
mecfs_status <- rbinom(50, 1, prob = 0.5)
niirf <- age + mecfs_status + rnorm(50, mean = 0, sd = 1)
model1 <- lm(niirf ~ 1) # Intercept only
model2 <- lm(niirf ~ mecfs_status)
anova_1_2 <- anova(model1, model2)
anova_1_2_pval <- anova_1_2[['Pr(>F)']][[2]]
model3 <- lm(niirf ~ age)
model4 <- lm(niirf ~ mecfs_status + age)
anova_3_4 <- anova(model3, model4)
anova_3_4_pval <- anova_3_4[['Pr(>F)']][[2]]
if (anova_1_2_pval > anova_3_4_pval) {
more_significant_count <- more_significant_count + 1
}
}
print(paste0("In ", more_significant_count, " out of ", total_trials, " total trials (", 100*more_significant_count/total_trials, "%), the p-value when adding ME/CFS status to an intercept-only model was higher (less significant) than the p-value when adding ME/CFS status to a model with an age covariate."))
anova_1_2
anova_3_4Output:
In 978 out of 1000 total trials (97.8%), the p-value when adding ME/CFS status to an intercept-only model was higher (less significant) than the p-value when adding ME/CFS status to a model with an age covariate.
To get an idea of what the p-values are, I looked at the ANOVA results for the last trial:
Code:
> anova_1_2
Analysis of Variance Table
Model 1: niirf ~ 1
Model 2: niirf ~ mecfs_status
Res.Df RSS Df Sum of Sq F Pr(>F)
1 49 3947.8
2 48 3906.9 1 40.879 0.5022 0.4819
> anova_3_4
Analysis of Variance Table
Model 1: niirf ~ age
Model 2: niirf ~ mecfs_status + age
Res.Df RSS Df Sum of Sq F Pr(>F)
1 48 52.230
2 47 47.129 1 5.1011 5.0872 0.0288 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1ScoutB
Senior Member (Voting Rights)
This explanation of restricted fraction and hindered fraction is exactly my understanding as well.The restricted fraction is the water inside small confined structures such as cells. It is water that is isotropic - it can move in all different directions, just not very far.
The hindered fraction is also isotropic, but I think excludes the restricted fraction and the FF. The NII-HR, the hindered water ratio is the ratio of that to the rest of the signal. I think it might be to the rest of the isotropic signal.
This was mentioned earlier but I think their "FF" here is fiber rather than free. Unlike other authors using the same techniques, this thread's paper does not seem to extract/calculate a value they are calling "free", (representing water that can move in all directions even more freely than the hindered and restricted fractions, equivalent to that far right red curve on the graph Hutan posted). I saw one author mention that this "free" fraction would be seen in big open cavities, like the ventricles.
I think the anisotropy is supposed to have been "removed" from the hindered and restricted components.I would expect water both inside and outside cells in white matter to have a degree of anisotropy to its diffusivity, certainly for outside and for nerve fibres. FF refers to fibres I think.
Is it that the anisotropy measures stratify across the 'hindered' and 'restricted' components?
My loose understanding of the math, incase it's of interest to anyone:
I think the idea is that they are trying to mathematically separate the isotropic and anisotropic parts of the diffusion. For each voxel, the MRI is able to tell them how freely the water can move in any direction they check. Then they model (with the equation below) how this MRI signal (S_k) breaks down into anisotropic (the sum on the left) and isotropic (the integral on the right) parts. They solve the set of formulas with numerical methods (I think the 'f's, including the function f(D) are the unknowns) and then (very roughly speaking) I think those 'f's are used to define the NII-RF, NII-HR values etc.
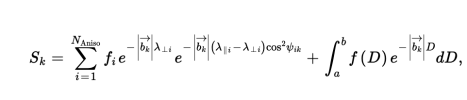
So the upshot is, I believe NII-RF and NII-HR are defined just using the parts of the signal they think are isotropic, according to their model.
Jonathan Edwards
Senior Member (Voting Rights)
I don't see our pathology ending up being lateralised like this.
Yes, this seems peculiar, for an immunopathology in particular.
jnmaciuch
Senior Member (Voting Rights)
Your intercept only comparison is not correct:Here is some R code simulating this with random data where NII-RF is correlated to age and to ME/CFS status:
Code:total_trials <- 1000 more_significant_count <- 0 for (i in 1:total_trials) { age <- rnorm(50, mean = 45, sd = 10) mecfs_status <- rbinom(50, 1, prob = 0.5) niirf <- age + mecfs_status + rnorm(50, mean = 0, sd = 1) model1 <- lm(niirf ~ 1) # Intercept only model2 <- lm(niirf ~ mecfs_status) anova_1_2 <- anova(model1, model2) anova_1_2_pval <- anova_1_2[['Pr(>F)']][[2]] model3 <- lm(niirf ~ age) model4 <- lm(niirf ~ mecfs_status + age) anova_3_4 <- anova(model3, model4) anova_3_4_pval <- anova_3_4[['Pr(>F)']][[2]] if (anova_1_2_pval > anova_3_4_pval) { more_significant_count <- more_significant_count + 1 } } print(paste0("In ", more_significant_count, " out of ", total_trials, " total trials (", 100*more_significant_count/total_trials, "%), the p-value when adding ME/CFS status to an intercept-only model was higher (less significant) than the p-value when adding ME/CFS status to a model with an age covariate.")) anova_1_2 anova_3_4
Output:
To get an idea of what the p-values are, I looked at the ANOVA results for the last trial:
When comparing the models without age, the p-value was 0.48. With age, it was 0.03.Code:> anova_1_2 Analysis of Variance Table Model 1: niirf ~ 1 Model 2: niirf ~ mecfs_status Res.Df RSS Df Sum of Sq F Pr(>F) 1 49 3947.8 2 48 3906.9 1 40.879 0.5022 0.4819 > anova_3_4 Analysis of Variance Table Model 1: niirf ~ age Model 2: niirf ~ mecfs_status + age Res.Df RSS Df Sum of Sq F Pr(>F) 1 48 52.230 2 47 47.129 1 5.1011 5.0872 0.0288 * --- Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Result:total_trials <- 1000
more_significant_count <- 0
for (i in 1:total_trials) {
age <- rnorm(50, mean = 45, sd = 10)
mecfs_status <- rbinom(50, 1, prob = 0.5)
niirf <- age + mecfs_status + rnorm(50, mean = 0, sd = 1)
model1 <- lm(niirf ~ 0) # Intercept only
model2 <- lm(niirf ~ 0 + mecfs_status)
anova_1_2 <- anova(model1, model2)
anova_1_2_pval <- anova_1_2[['Pr(>F)']][[2]]
model3 <- lm(niirf ~ age)
model4 <- lm(niirf ~ mecfs_status + age)
anova_3_4 <- anova(model3, model4)
anova_3_4_pval <- anova_3_4[['Pr(>F)']][[2]]
if (anova_1_2_pval > anova_3_4_pval) {
more_significant_count <- more_significant_count + 1
}
}
print(paste0("In ", more_significant_count, " out of ", total_trials, " total trials (", 100*more_significant_count/total_trials, "%), the p-value when adding ME/CFS status to an intercept-only model was higher (less significant) than the p-value when adding ME/CFS status to a model with an age covariate."))
anova_1_2
anova_3_4
"In 13 out of 1000 total trials (1.3%), the p-value when adding ME/CFS status to an intercept-only model was higher (less significant) than the p-value when adding ME/CFS status to a model with an age covariate."
or, allowing intercept fitting (the better comparison):
Result:for (i in 1:total_trials) {
age <- rnorm(50, mean = 45, sd = 10)
mecfs_status <- rbinom(50, 1, prob = 0.5)
niirf <- age + mecfs_status + rnorm(50, mean = 0, sd = 1)
model1 <- lm(niirf ~ 0) # Intercept only
model2 <- lm(niirf ~ mecfs_status)
anova_1_2 <- anova(model1, model2)
anova_1_2_pval <- anova_1_2[['Pr(>F)']][[2]]
model3 <- lm(niirf ~ age)
model4 <- lm(niirf ~ mecfs_status + age)
anova_3_4 <- anova(model3, model4)
anova_3_4_pval <- anova_3_4[['Pr(>F)']][[2]]
if (anova_1_2_pval > anova_3_4_pval) {
more_significant_count <- more_significant_count + 1
}
}
"In 0 out of 1000 total trials (0%), the p-value when adding ME/CFS status to an intercept-only model was higher (less significant) than the p-value when adding ME/CFS status to a model with an age covariate."
[note: sorry for all the editing, was writing quickly. I don't think the intercept is the major problem here--I figured it would be faster to write code that tests the issue I brought up directly (see post #86)]
Last edited:
Jonathan Edwards
Senior Member (Voting Rights)
So the upshot is, I believe NII-RF and NII-HR are defined just using the parts of the signal they think are isotropic, according to their model.
That sounds right, although of course in neuroinflammation I suspect there will be a degree of anisoprotpy of water diffusion in white matter even for extracellular free water because of the linear architecture.
The separation of these various different parameters seems very clever. What I am a bit dubious about its the interpretation. Especially when looking at the colour pictures, as SNTG says.
Your intercept only comparison is not correct:
You replaced
model1 <- lm(niirf ~ 1) with model1 <- lm(niirf ~ 0). 1 is the formula for an intercept. 0 means no intercept. From the R docs:
A formula has an implied intercept term. To remove this use either y ~ x - 1 or y ~ 0 + x.
So in the first updated code you posted, in the comparison of model1 and model2, it's comparing a model that just predicts 0 for every point with a model that predicts based on mecfs_status, but is forced through the point (0,0).
In the second code, it's comparing a model which predicts 0 for every point to a model that includes mecfs_status and an intercept.
jnmaciuch
Senior Member (Voting Rights)
Sorry, short on time today otherwise would write more. [edit the important thing] to check is that the case I'm talking about is where the original NII-XX ~ ME/CFS association wasn't significant initially. Even if you take mecfs_status out of the outcome variable, you might still generate a spurious association randomly. The issue I've been talking about is the measurements besides NII-RF that dropped out in the univariateYou replacedmodel1 <- lm(niirf ~ 1)withmodel1 <- lm(niirf ~ 0). 1 is the formula for an intercept. 0 means no intercept.
From the R docs:
So in the first updated code you posted, in the comparison of model1 and model2, it's comparing a model that just predicts 0 for every point with a model that predicts based on mecfs_status, but is forced through the point (0,0).
In the second code, it's comparing a model which predicts 0 for every point to a model that includes mecfs_status and an intercept.
Last edited:
jnmaciuch
Senior Member (Voting Rights)
Okay here is code testing this specific scenario:
The situation I flagged, where a variable was not significant in a univariate association but is significant with age as a covariate, only occured 5% of the time, so exactly what was expected by chance. Meaning that a confounder that is associated with the outcome variable does not induce significance in the association between ME/CFS and the outcome var if included as a covariate. Meanwhile, in the paper, 6/7 measured NII variables showed this trend.
@forestglip let me know if this makes sense
I'm not sure about that.
Say you have a feature that increases a lot by age but is reliably lower in ME/CFS at each age. If you had a badly matched control group, a lot younger, then there might not be a significant difference between the ME/CFS group and the control group for that feature. So, there would be no obvious association to start with. But then, if you did control for age, the difference would appear.
library(dplyr)
library(car)
total_trials <- 5000
null_univariate <- 0
null_univariate_sig_multivariate <- 0
for (i in 1:total_trials) {
age <- rnorm(50, mean = 45, sd = 20)
mecfs_status <- rbinom(50, 1, prob = 0.5)
# Create variable that is associated with age
niiXX <- age + rnorm(50, mean = 0, sd = 1)
# Check that mecfs_status is NOT significantly associated with outcome on its own
model1 <- lm(niiXX ~ mecfs_status)
pval <- car::Anova(model1)$`Pr(>F)`[1]
# If univariate association is not significant
if (pval > 0.05) {
null_univariate <- null_univariate + 1
# Determine if mecfs_status is significant in a model with age as a covariate
model2 <- lm(niiXX ~ mecfs_status + age)
pval_multi <- car::Anova(model2)$`Pr(>F)`[1]
if (pval_multi < 0.05) {
null_univariate_sig_multivariate <- null_univariate_sig_multivariate + 1
}
}
}
print(paste0("In ", null_univariate_sig_multivariate, " out of ", null_univariate, " total trials (", signif(100*null_univariate_sig_multivariate/null_univariate, 2), "%), the ME/CFS variable with significant with the covariate of age but not on its own."))
library(car)
total_trials <- 5000
null_univariate <- 0
null_univariate_sig_multivariate <- 0
for (i in 1:total_trials) {
age <- rnorm(50, mean = 45, sd = 20)
mecfs_status <- rbinom(50, 1, prob = 0.5)
# Create variable that is associated with age
niiXX <- age + rnorm(50, mean = 0, sd = 1)
# Check that mecfs_status is NOT significantly associated with outcome on its own
model1 <- lm(niiXX ~ mecfs_status)
pval <- car::Anova(model1)$`Pr(>F)`[1]
# If univariate association is not significant
if (pval > 0.05) {
null_univariate <- null_univariate + 1
# Determine if mecfs_status is significant in a model with age as a covariate
model2 <- lm(niiXX ~ mecfs_status + age)
pval_multi <- car::Anova(model2)$`Pr(>F)`[1]
if (pval_multi < 0.05) {
null_univariate_sig_multivariate <- null_univariate_sig_multivariate + 1
}
}
}
print(paste0("In ", null_univariate_sig_multivariate, " out of ", null_univariate, " total trials (", signif(100*null_univariate_sig_multivariate/null_univariate, 2), "%), the ME/CFS variable with significant with the covariate of age but not on its own."))
I randomly generated values for an outcome variable (niiXX) that was deliberately coded to correlate with age. I then checked the association between niiXX and mecfs_status. If that univariate association was not significant (p > 0.05), I tested a second model including mecfs_status and age.[1] "In 243 out of 4721 total trials (5.1%), the ME/CFS variable with significant with the covariate of age but not on its own."
The situation I flagged, where a variable was not significant in a univariate association but is significant with age as a covariate, only occured 5% of the time, so exactly what was expected by chance. Meaning that a confounder that is associated with the outcome variable does not induce significance in the association between ME/CFS and the outcome var if included as a covariate. Meanwhile, in the paper, 6/7 measured NII variables showed this trend.
@forestglip let me know if this makes sense
Last edited:
This is because in the model you made, there is no relationship between ME/CFS and niXX, so it is all due to chance. niXX is just a function of age, so there should only be about 5% significant as false positives when testing the association with mecfs_status.I randomly generated values for an outcome variable (niiXX) that was deliberately coded to correlate with age. I then checked the association between niiXX and mecfs_status. If that univariate association was not significant (p > 0.05), I tested a second model including mecfs_status and age.
The situation I flagged, where a variable was not significant in a univariate association but is significant with covariates, only occured 5% of the time, so exactly what was expected by chance.
If niXX actually depends on mecfs_status, then adding the covariate can make it more significant, and can take p from greater than 0.05 to less than 0.05. I added mecfs_status to the initial niXX model in your code:
niiXX <- age + mecfs_status + rnorm(50, mean = 0, sd = 1)
Resulting in:
In 4345 out of 4731 total trials (92%), the ME/CFS variable with significant with the covariate of age but not on its own.
