library(tidyverse)
library(magrittr)
library(data.table)
library(colourvalues)
# Load summary stats
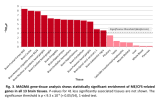
# Inputs are 6 csv files manually exported from jumping to top loci at
LocusZoom
files <- list.files(workDir,
pattern = ".csv")
regions <- files %>%
str_split_i(pattern = "_",
i = 2)
summary_stats <- files %>%
map(\(x) read_csv(x)) %>%
set_names(regions)
# Filter to credible set
summary_stats %<>%
map(\(x) x %>%
as.data.table() %>%
.[`Cred. set` == TRUE])
# Bind rows
summary_stats %<>% rbindlist(idcol = "region")
# Rename some columns
summary_stats %<>% setnames(old = c("Chrom", "Pos", "-log<sub>10</sub>(p)", "β", "Alt freq."),
new = c("chrom", "chromEnd", "neglog10pval", "beta", "alt_freq"))
# Format chrom column
summary_stats %<>% .[, chrom := paste0("chr", chrom)]
# Add start position
summary_stats %<>% .[, chromStart := chromEnd - 1]
# Create name
summary_stats %<>% .[, name := paste0(Ref, ">", Alt)]
# Create description
summary_stats %<>% .[, desc := paste0("rsID=",
rsID,
"; Beta=",
signif(beta, 4),
"; -log10(pval)=",
signif(neglog10pval, 3),
"; alt freq=",
alt_freq)]
# Around each peak, scale color values into 8 bins (8 is color limit) and assign RGB code
summary_stats %<>% .[, color_bin := cut_number(neglog10pval, n = 8) %>%
as.numeric(),
by = "region"] %>%
.[, color := colour_values_rgb(color_bin,
include_alpha = F) %>%
apply(MARGIN = 1, \(x) paste(x, collapse = ","))]
# Add ID
summary_stats %<>% .[, ID := 1:nrow(.)]
# Add score
summary_stats %<>% .[, score := 999] %>%
.[, strand := "+"]
# Pull columns for BED file
BED <- summary_stats %>%
.[, c("chrom",
"chromStart",
"chromEnd",
"name",
"score",
"strand",
"chromStart",
"chromEnd",
"color",
"ID",
"desc")]
# Add track header as column names
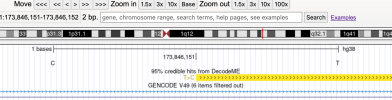
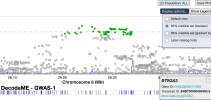
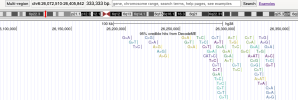
header <- c("track name=DecodeME",
"type='bedDetail'",
"description='95% credible hits from DecodeME'",
"db=hg38",
"visibility=3",
"itemRgb='On'",
"",
"",
"",
"",
"")
BED %<>% setnames(header)
# Save as tab delimited file
write_delim(BED,
file = file.path(workDir,
"DecodeME_CredibleSet_UCSC_custom_track.txt"),
delim = "\t")