Obermann
Senior Member (Voting Rights)
Abstract
Background: Neurologic Post Treatment Lyme disease (nPTLS) and Chronic Fatigue (CFS) are syndromes of unknown etiology. They share features of fatigue and cognitive dysfunction, making it difficult to differentiate them. Unresolved is whether nPTLS is a subset of CFS.
Methods and principal findings: Pooled cerebrospinal fluid (CSF) samples from nPTLS patients, CFS patients, and healthy volunteers were comprehensively analyzed using high-resolution mass spectrometry (MS), coupled with immunoaffinity depletion methods to reduce protein-masking by abundant proteins. Individual patient and healthy control CSF samples were analyzed directly employing a MS-based label-free quantitative proteomics approach. We found that both groups, and individuals within the groups, could be distinguished from each other and normals based on their specific CSF proteins (p<0.01). CFS (n = 43) had 2,783 non-redundant proteins, nPTLS (n = 25) contained 2,768 proteins, and healthy normals had 2,630 proteins. Preliminary pathway analysis demonstrated that the data could be useful for hypothesis generation on the pathogenetic mechanisms underlying these two related syndromes.
Conclusions: nPTLS and CFS have distinguishing CSF protein complements. Each condition has a number of CSF proteins that can be useful in providing candidates for future validation studies and insights on the respective mechanisms of pathogenesis. Distinguishing nPTLS and CFS permits more focused study of each condition, and can lead to novel diagnostics and therapeutic interventions.
How strong is the Schutzer et al. study from 2011?
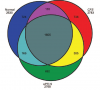
I have always thought that the proteomic study on the cerebrospinal fluid by Schutzer et al in 2011 was rock solid. It was selected as one of the top 100 scientific studies in 2011 by Discover Magazine. The Venn diagram (above), where they showed many unique proteins for patients with ME/CFS, was impressive. I just got into an argument with an ME-sceptic professor about that study, and she said that the differences in the Venn diagram weren't significant. After reviewing the supplementary table S1 in the paper, I have to say that I agree. Many proteins that were listed as unique for ME/CFS were positive only for one patient.
Any thoughts on that? @Jonathan Edwards ?
Edit: The samples were pooled, so we don't know how many patients in each group who had the proteins. Table S1 lists the number of unique peptides that were found. However, for the proteins that were unique for a study group, usually only one unique peptide was identified; whereas several tens or hundreds of unique peptides were found for proteins that were detected in all study groups. That does suggest that only few individuals had unique proteins.
Background: Neurologic Post Treatment Lyme disease (nPTLS) and Chronic Fatigue (CFS) are syndromes of unknown etiology. They share features of fatigue and cognitive dysfunction, making it difficult to differentiate them. Unresolved is whether nPTLS is a subset of CFS.
Methods and principal findings: Pooled cerebrospinal fluid (CSF) samples from nPTLS patients, CFS patients, and healthy volunteers were comprehensively analyzed using high-resolution mass spectrometry (MS), coupled with immunoaffinity depletion methods to reduce protein-masking by abundant proteins. Individual patient and healthy control CSF samples were analyzed directly employing a MS-based label-free quantitative proteomics approach. We found that both groups, and individuals within the groups, could be distinguished from each other and normals based on their specific CSF proteins (p<0.01). CFS (n = 43) had 2,783 non-redundant proteins, nPTLS (n = 25) contained 2,768 proteins, and healthy normals had 2,630 proteins. Preliminary pathway analysis demonstrated that the data could be useful for hypothesis generation on the pathogenetic mechanisms underlying these two related syndromes.
Conclusions: nPTLS and CFS have distinguishing CSF protein complements. Each condition has a number of CSF proteins that can be useful in providing candidates for future validation studies and insights on the respective mechanisms of pathogenesis. Distinguishing nPTLS and CFS permits more focused study of each condition, and can lead to novel diagnostics and therapeutic interventions.
How strong is the Schutzer et al. study from 2011?

I have always thought that the proteomic study on the cerebrospinal fluid by Schutzer et al in 2011 was rock solid. It was selected as one of the top 100 scientific studies in 2011 by Discover Magazine. The Venn diagram (above), where they showed many unique proteins for patients with ME/CFS, was impressive. I just got into an argument with an ME-sceptic professor about that study, and she said that the differences in the Venn diagram weren't significant. After reviewing the supplementary table S1 in the paper, I have to say that I agree. Many proteins that were listed as unique for ME/CFS were positive only for one patient.
Any thoughts on that? @Jonathan Edwards ?
Edit: The samples were pooled, so we don't know how many patients in each group who had the proteins. Table S1 lists the number of unique peptides that were found. However, for the proteins that were unique for a study group, usually only one unique peptide was identified; whereas several tens or hundreds of unique peptides were found for proteins that were detected in all study groups. That does suggest that only few individuals had unique proteins.
Last edited by a moderator:
