Simon M
Senior Member (Voting Rights)
DNA sequencing study to help pinpoint biology of ME gets £4.7m
The science of Sequence ME and Long Covid
The UK government has awarded nearly £5 million to fund full sequencing of the DNA of 6,000 people with ME using the best available technology. This will help scientists home in on what is really driving the disease. And will probably be the biggest study of its kind for any disease.
The UK government has committed £4.7 million directly to ME/CFS research through an award from its Office for Life Sciences to the landmark study Sequence ME & Long Covid. The award will pay for full sequencing of the DNA of 6,000 people with ME/CFS, using the method of British biotech company Oxford Nanopore Technologies.
This is an extraordinary moment for ME/CFS. It used to get by on the dregs of research funding, but now has a study that will probably be the largest of any single disease using this technique. It Is the result of a lot of work by a lot of people.
The DNA samples are already available through the DecodeME study. Sequence ME & Long Covid ultimately aims to sequence 9,000 people with ME/CFS and 9,000 with Long Covid – as funds permit. The project is led by the University of Edinburgh and Action for ME.
Full sequencing can reveal more
The new method gives real depth of genetic detail. DecodeME used older, much cheaper technology looking at common DNA differences at about half a million places in the whole genome. It found a genetic signal in eight small regions of DNA, but these contained dozens of genes, and it isn’t yet clear which ones are involved in ME/CFS.
Using the Oxford Nanopore Technologies approach, Sequence ME & Long Covid will be able to sequence almost all of the 3 billion chemical 'letters' in the human genome. As Professor Chris Ponting, scientific lead for the study said,
“This project will allow us to pinpoint individual genes disrupted in ME/CFS… breaking down this complex disease into its underlying biological causes – bringing us closer to more precise diagnosis and, ultimately, targeted treatments.”
The technology
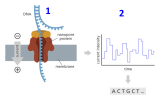
At its heart, the sophisticated Oxford Nanopore technology is simple, as shown in the image above,
30 second video
read the full blog for more:
The science of Sequence ME and Long Covid
The UK government has awarded nearly £5 million to fund full sequencing of the DNA of 6,000 people with ME using the best available technology. This will help scientists home in on what is really driving the disease. And will probably be the biggest study of its kind for any disease.
The UK government has committed £4.7 million directly to ME/CFS research through an award from its Office for Life Sciences to the landmark study Sequence ME & Long Covid. The award will pay for full sequencing of the DNA of 6,000 people with ME/CFS, using the method of British biotech company Oxford Nanopore Technologies.
This is an extraordinary moment for ME/CFS. It used to get by on the dregs of research funding, but now has a study that will probably be the largest of any single disease using this technique. It Is the result of a lot of work by a lot of people.
The DNA samples are already available through the DecodeME study. Sequence ME & Long Covid ultimately aims to sequence 9,000 people with ME/CFS and 9,000 with Long Covid – as funds permit. The project is led by the University of Edinburgh and Action for ME.
Full sequencing can reveal more
The new method gives real depth of genetic detail. DecodeME used older, much cheaper technology looking at common DNA differences at about half a million places in the whole genome. It found a genetic signal in eight small regions of DNA, but these contained dozens of genes, and it isn’t yet clear which ones are involved in ME/CFS.
Using the Oxford Nanopore Technologies approach, Sequence ME & Long Covid will be able to sequence almost all of the 3 billion chemical 'letters' in the human genome. As Professor Chris Ponting, scientific lead for the study said,
“This project will allow us to pinpoint individual genes disrupted in ME/CFS… breaking down this complex disease into its underlying biological causes – bringing us closer to more precise diagnosis and, ultimately, targeted treatments.”
The technology

At its heart, the sophisticated Oxford Nanopore technology is simple, as shown in the image above,
- A strand of DNA is threaded through a tiny hole in a membrane - a nanopore.
- Each of the four chemical letters (A, T, G and C) creates a distinct electrical signal as it passes through the pore – and the DNA sequence can be read out from the electrical trace, one letter at a time.
30 second video
read the full blog for more:
- why full sequencing help pinpoints the biology behind ME/CFS
- why the long read of the nanopore approach is so important (finds genetic changes other methods miss
- plus nanopore method detects gene silencing (epigenetics)
- cutting edge data analysis from the European Bioinformatics Institute, a partner in the study
Last edited:
