You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
Biological Insights from Genome-Wide Association Studies and Whole Genome Sequencing of [ME/CFS], 2026, Maccallini et al
- Thread starter Amw66
- Start date
Jonathan Edwards
Senior Member (Voting Rights)
Is he the 'citizen scientist' Chris P referred to?
Biological Insights from Genome-Wide Association Studies and Whole Genome Sequencing of Myalgic Encephalomyelitis/Chronic Fatigue Syndrome
Maccallini, Paolo
Abstract
Myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) is a debilitating disorder of poorly understood etiology.
We performed a meta-analysis of two European-ancestry ME/CFS genome-wide association studies (GWAS) with no overlap in subjects, DecodeME and Million Veteran Program, comprising a total of 19,470 cases and 699,111 controls. Post-GWAS analyses investigated the association between ME/CFS and specific tissues, cell types, cellular components, and canonical pathways. Findings were independently evaluated for replication against a module of ME/CFS risk genes previously prioritized by machine learning applied to rare coding variants from whole-genome sequencing (WGS) of ME/CFS cases and controls of European ancestry.
Tissue enrichment analysis implicated multiple brain regions and the pituitary, with no peripheral tissue reaching significance, and three survived Bonferroni-corrected replication. Gene-set analysis identified multiple neuronal and synaptic gene sets, several of which were independently replicated, with glutamatergic synapses as the most specific replicated signal. Cell-type analysis identified independent replicated signals in distinct neuronal populations of subcortical and cerebellar regions.
These results suggest a role for synaptic function in specific brain regions in the pathogenesis of ME/CFS, with convergent support from both common and rare variant data. Larger studies are needed to confirm these findings and to identify specific targets for therapeutic intervention.
Web | DOI | Zenodo | Preprint
Maccallini, Paolo
Abstract
Myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) is a debilitating disorder of poorly understood etiology.
We performed a meta-analysis of two European-ancestry ME/CFS genome-wide association studies (GWAS) with no overlap in subjects, DecodeME and Million Veteran Program, comprising a total of 19,470 cases and 699,111 controls. Post-GWAS analyses investigated the association between ME/CFS and specific tissues, cell types, cellular components, and canonical pathways. Findings were independently evaluated for replication against a module of ME/CFS risk genes previously prioritized by machine learning applied to rare coding variants from whole-genome sequencing (WGS) of ME/CFS cases and controls of European ancestry.
Tissue enrichment analysis implicated multiple brain regions and the pituitary, with no peripheral tissue reaching significance, and three survived Bonferroni-corrected replication. Gene-set analysis identified multiple neuronal and synaptic gene sets, several of which were independently replicated, with glutamatergic synapses as the most specific replicated signal. Cell-type analysis identified independent replicated signals in distinct neuronal populations of subcortical and cerebellar regions.
These results suggest a role for synaptic function in specific brain regions in the pathogenesis of ME/CFS, with convergent support from both common and rare variant data. Larger studies are needed to confirm these findings and to identify specific targets for therapeutic intervention.
Web | DOI | Zenodo | Preprint
Jonathan Edwards
Senior Member (Voting Rights)
Were you aware of this one @James Cox ?
ME/CFS Science Blog
Senior Member (Voting Rights)
This is impressive work and well worth an in depth-read!
It follows the same line of reasoning we've been talking about on the forum fore more than a year, based on both the DecodeME data on common mutations and Mark Synder study on rare mutations hinting at neural synapses in the pathology of ME/CFS. But Paolo has done it in a more formal and persuasive way.
A first important addition is that he managed to add the 3891 ME/CFS cases from the Million Veteran Program (MVP) to the 15,579 from DecodeME in a meta-analysis. Paolo said he wanted to add cases from other genetic databases as well such as the UK biobank or FinnGenn, but that gave the issue that some of the controls overlapped. So this (DecodeME + MVP) is the biggest meta-analysis he could do.
Those from MVP were based on ICD-codes and with a prevalences of around 1.5% they might contain a lot of false positives (patients label ME/CFS who might not have it). But the bump in statistical power was just enough to get a lot of interesting results.
Paolo used the MAGMA tool that uses database on gene expression to see if the DNA results from ME/CFS patients are enriched in a particular group of genes or tissue. The DecodeME preprint did this as well and we (mostly forestglip) played around with this on the forum as well, showing it mostly pointed to the brain. But with the added MVP samples, Paolo seem to have gotten much stronger result implicating synapses in multiple ways.

The second interesting thing he added is that he used these gene sets from MAGMA and tested them in the Snyder study that used rare mutations. Several of these gene sets were replicated. Here are the important paragraphs that describe this approach:
The tissue results were similar to those from DecodeME alone I believe (various brain regions), but using various gene expression databases, Paolo found significant hits for particular brain cell types, most notably:
- the eccentric medium spiny neuron (eMSN) in the claustrum
- glutamatergic neuron in cerebellar white matter
It follows the same line of reasoning we've been talking about on the forum fore more than a year, based on both the DecodeME data on common mutations and Mark Synder study on rare mutations hinting at neural synapses in the pathology of ME/CFS. But Paolo has done it in a more formal and persuasive way.
A first important addition is that he managed to add the 3891 ME/CFS cases from the Million Veteran Program (MVP) to the 15,579 from DecodeME in a meta-analysis. Paolo said he wanted to add cases from other genetic databases as well such as the UK biobank or FinnGenn, but that gave the issue that some of the controls overlapped. So this (DecodeME + MVP) is the biggest meta-analysis he could do.
Those from MVP were based on ICD-codes and with a prevalences of around 1.5% they might contain a lot of false positives (patients label ME/CFS who might not have it). But the bump in statistical power was just enough to get a lot of interesting results.
Paolo used the MAGMA tool that uses database on gene expression to see if the DNA results from ME/CFS patients are enriched in a particular group of genes or tissue. The DecodeME preprint did this as well and we (mostly forestglip) played around with this on the forum as well, showing it mostly pointed to the brain. But with the added MVP samples, Paolo seem to have gotten much stronger result implicating synapses in multiple ways.

The second interesting thing he added is that he used these gene sets from MAGMA and tested them in the Snyder study that used rare mutations. Several of these gene sets were replicated. Here are the important paragraphs that describe this approach:
Gene-set analysis using MAGMA tested 17,009 of the 17,023 gene sets in the input file(MSigDB_20231Hs_MAGMA.txt). Twelve gene sets were significantly enriched for association signalin the DME-1 + MVP meta-analysis after Benjamini-Hochberg correction (PBH < 0.05, k = 17,009;Table 4). All 12 span neuronal and synaptic categories from Gene Ontology cellular components,Gene Ontology molecular functions, and Reactome. Three of the 12 also survived the more stringent Bonferroni correction (P < 0.05/17,009 = 2.94 × 10−6): postsynaptic specialization, neuron-to-neuronsynapse, and somatodendritic compartment, all Gene Ontology cellular component terms implicating the synapse in general and post-synapses in particular. Neither DME-1 nor MVP alone reached significant enrichment after BH correction (S1 File, sheet S3).
I think this is quite big, because it's not just about DecodeME (one study anymore) or even one approach (GWAS of common alleles).Replication was evaluated using the HEAL2 module of ME/CFS risk genes [20]. Only 6,063 of the17,023 input gene sets had non-empty overlap with the HEAL2 background (17,759 genes) and were testable. Seven of the 12 BH-significant gene sets were replicated at Bonferroni-corrected significance(P < 0.05/12 = 4.17 × 10−3; shaded rows in Table 4), of which six also survived Bonferroni correctionin the HEAL2 module (P < 0.05/6,063 = 8.25 × 10−6; bold replication p-values in Table 4). All three gene sets surviving Bonferroni correction in discovery were replicated at Bonferroni-correctedsignificance (P < 0.05/3 = 0.017; P†in Table 4).
The tissue results were similar to those from DecodeME alone I believe (various brain regions), but using various gene expression databases, Paolo found significant hits for particular brain cell types, most notably:
- the eccentric medium spiny neuron (eMSN) in the claustrum
- glutamatergic neuron in cerebellar white matter
Last edited:
ME/CFS Science Blog
Senior Member (Voting Rights)
These were the 5 hits in the meta-analysis. The one on chromosome 17 for CA10 is not there but we do have a new one on chromosome 19.
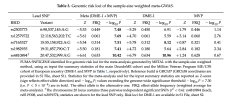
Paolo provided the GRCh38 location in the supplementary material:
When I looked up the location on chr 19 on LocusZoom I got the following picture:

TSHZ3 seems to be involved in neural development.

Paolo provided the GRCh38 location in the supplementary material:
| topLeadSNP | GRCh38 | GRCh37 |
| rs2503773 | 6:98089269:A:G | 6:98537145:A:G |
| rs12579722 | 12:118125397:T:C | 12:118563202:T:C |
| rs7165327 | 15:54866724:A:G | 15:55158922:A:G |
| rs1982935 | 19:31366884:C:T | 19:31857790:C:T |
| rs4810894 | 20:48916462:A:G | 20:47532999:A:G |
When I looked up the location on chr 19 on LocusZoom I got the following picture:

TSHZ3 seems to be involved in neural development.
Jonathan Edwards
Senior Member (Voting Rights)
the eccentric medium spiny neuron (eMSN) in the claustrum
I wonder if people with ME/CFS want to be thought of as medium spiny and in the claustrum?
I must admit I had not thought of that!
